Specific Aims
The following outlines a few experiments that would ultimately measure gene expression of EGFR mutants mutant mice under the influence of different small molecules. Performing these methods on epithelial cells may uncover important cellular function underlying Lung Adenocarcinoma
Aim 1
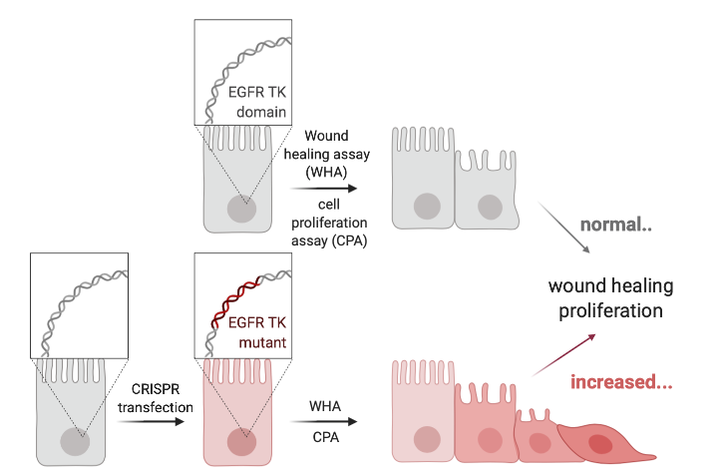
Determine which part of the tyrosine kinase domain in EGFR influences cell migration and proliferation.
Approach: Ensemble and ClustalOmega will be used to align known homologs of the EGFR gene and I will identify unique Amino acids in the Tyrosine kinase domain that might be important for cell proliferation and cell adhesion in the lung. CRISPR/Cas9 plasmid delivery to mouse eggs will mutate the corresponding DNA segments in mouse eggs7, followed by DNA sequencing of wildtype and transfected mice. To measure cell the rate of proliferation and migration in the lung, I would use a cell counting kit and a wound healing (scratch) assay, respectively.
Rationale: Identifying the exact amino acid in EGFR, that leads to changes in the rate of proliferation and migration, will be important for understanding how EGFR signals normally in the lung to mediate cell.
Hypothesis: I hypothesize that mutation of tyrosine kinase specific residues, specifically in exon 19 and 20 will result in an increase in cell proliferation and adhesion.
Approach: Ensemble and ClustalOmega will be used to align known homologs of the EGFR gene and I will identify unique Amino acids in the Tyrosine kinase domain that might be important for cell proliferation and cell adhesion in the lung. CRISPR/Cas9 plasmid delivery to mouse eggs will mutate the corresponding DNA segments in mouse eggs7, followed by DNA sequencing of wildtype and transfected mice. To measure cell the rate of proliferation and migration in the lung, I would use a cell counting kit and a wound healing (scratch) assay, respectively.
Rationale: Identifying the exact amino acid in EGFR, that leads to changes in the rate of proliferation and migration, will be important for understanding how EGFR signals normally in the lung to mediate cell.
Hypothesis: I hypothesize that mutation of tyrosine kinase specific residues, specifically in exon 19 and 20 will result in an increase in cell proliferation and adhesion.
Aim 2
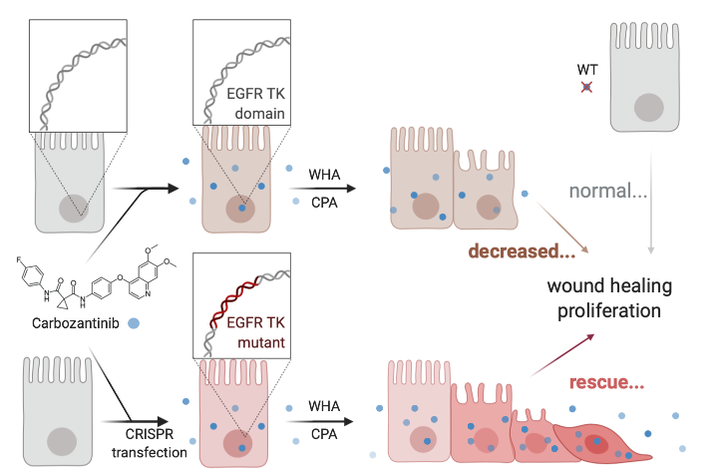
Determine how tyrosine-independent EGFR function influences cell proliferation and migration.
Approach: Reverse chemical genomic screening will quantify exogenous ligand to EGFR between wildtype and transfected mice. Samples will be subject to a variety of chemicals such as hydroxylamine that cleave cysteine- palmitoylation bonds8. Mutant epithelial and control sample tissues will be subjected to these chemicals. Once again, cultures will be measured for proliferation and cell migration by scratch assay and cell counting.
Rationale: Quantifying the effects of exogenous ligands on cell proliferation and migration between wild type and mutant mice will illustrate how prevalent tyrosine dependent function is for these phenotypes.
Hypothesis: I propose that chemical disruption of loss of function tyrosine kinase mutant cells will affect alternative mechanisms of increased cell proliferation and migration.
Approach: Reverse chemical genomic screening will quantify exogenous ligand to EGFR between wildtype and transfected mice. Samples will be subject to a variety of chemicals such as hydroxylamine that cleave cysteine- palmitoylation bonds8. Mutant epithelial and control sample tissues will be subjected to these chemicals. Once again, cultures will be measured for proliferation and cell migration by scratch assay and cell counting.
Rationale: Quantifying the effects of exogenous ligands on cell proliferation and migration between wild type and mutant mice will illustrate how prevalent tyrosine dependent function is for these phenotypes.
Hypothesis: I propose that chemical disruption of loss of function tyrosine kinase mutant cells will affect alternative mechanisms of increased cell proliferation and migration.
Aims 3
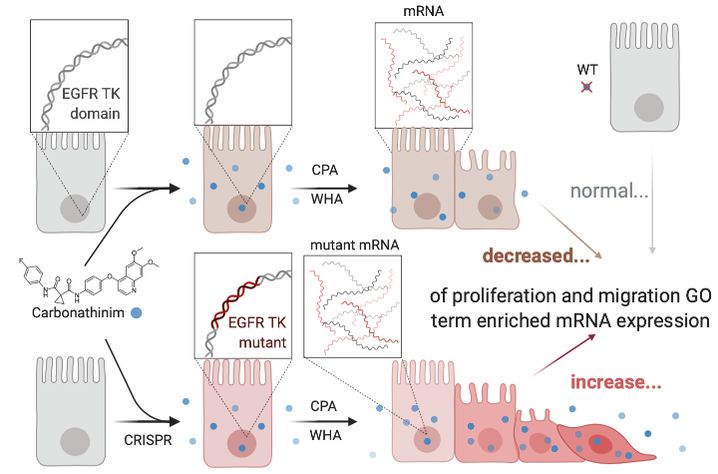
Identify how EGFR function affects gene expression associated with epithelial transformation.
Approach: I will perform a microarray on wildtype and transfected mice mouse cells treated with chemicals from the screen. Afterwards, Gene Ontology analysis using PANTHER can enrich gene expression with GO terms relating to cell growth patterns such as cell proliferation, migration and protein palmitoylation which are all implicated in epithelial transformations.
Rationale: Determining changes in proliferation, migration and protein palmitoylation related gene expression between cell samples will illustrate how EGFR binding mediates epithelial transformation.
Hypothesis: I hypothesize that mutant EGFR cell samples retains the ability to increase gene expression that can be affected by exogenous chemicals, through tyrosine kinase-independent mechanisms.
Approach: I will perform a microarray on wildtype and transfected mice mouse cells treated with chemicals from the screen. Afterwards, Gene Ontology analysis using PANTHER can enrich gene expression with GO terms relating to cell growth patterns such as cell proliferation, migration and protein palmitoylation which are all implicated in epithelial transformations.
Rationale: Determining changes in proliferation, migration and protein palmitoylation related gene expression between cell samples will illustrate how EGFR binding mediates epithelial transformation.
Hypothesis: I hypothesize that mutant EGFR cell samples retains the ability to increase gene expression that can be affected by exogenous chemicals, through tyrosine kinase-independent mechanisms.
Files
Specific Aims Draft 1:
| specific_aims_draft_1-kye_nichols.docx |
Specific Aims Draft 2:
| specific_aims_draft_2-kye_nichols.docx |
Specific Aims Final Draft:
| kyenicholsfinalaims2020.docx |
References
1.) Zhang, Natasha B. Leighl, et. al (2019). Emerging therapies for non-small cell lung cancer [Journal of Hematology and Oncology Article], pg 1. Retrieved from: https://jhoonline.biomedcentral.com/articles/10.1186/s13045-019-0731-8
2.) Kern, Tjaden, et al., Inhibitors of the epidermal growth factor receptor in apple juice extract. (2005). Retrieved from: https://www.ncbi.nlm.nih.gov/pubmed/15759309
3.) Sven Bogdan, Klämbt, Epidermal growth factor receptor signaling (2001), pg 3. Retrieved from: https://www.cell.com/current-biology/comments/S0960-9822(01)00167-1
4.) Thomas, Weihua, Rethink of EGFR in Cancer With Its Kinase Independent Function on Board (August, 2019). Retrieved from: https://www.frontiersin.org/articles/10.3389/fonc.2019.00800/full
5.) Tian, Shi, et. al (2018). Different subtypes of EGFR exon19 mutation can affect prognosis of patients with non-small cell lung adenocarcinoma [Pubmed Article]. Retrieved from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6211626/
6.) Tabeta, Hoebe, et al, Velvet, a Dominant Egfr Mutation That Causes Wavy Hair and Defective Eyelid Development in Mice (2003). Retrieved from: https://pdfs.semanticscholar.org/7eee/c89406e69357fa8ccf3874696448de616699.pdf?_ga=2.21516347.1433771477.1588236341-1015871252.1588236341
7.) Ohashi, Rai, et al., Induction of lung adenocarcinoma in transgenic mice expressing activated EGFR driven by the SP‐C promoter (2008). Retrieved from: https://www.ncbi.nlm.nih.gov/pubmed/18564139
8.) Martin, Chemical approaches for profiling dynamic palmitoylation (2013). Retrieved from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3693551/
9.) Cancer Treatment Centers of America, Lung cancer types, pg 2. Retrieved from: https://www.cancercenter.com/cancer-types/lung-cancer/types
10.) Uramoto, Iwata, et. al (2010). Epithelial–Mesenchymal Transition in EGFR-TKI Acquired Resistant Lung Adenocarcinoma [Pubmed Article]. Retrieved from: https://www.ncbi.nlm.nih.gov/pubmed/20682976
11.) Sawabata, Circulating tumor cells in lung cancer: cluster circulating tumor cells as hybrid epithelial-mesenchymal transition/mesenchymal-epithelial transition (2017). Retrieved from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5723813/
12.) Lin, Wang, et al., EGFR-TKI resistance in NSCLC patients: mechanisms and strategies (2014). Retrieved from: https://www.nature.com/articles/onc2012158
13.) Singh, Behrens, et al., Phosphorylation of MUC1 by Met Modulates Interaction with p53 and MMP1 Expression (2008). Retrieved from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4163608/
2.) Kern, Tjaden, et al., Inhibitors of the epidermal growth factor receptor in apple juice extract. (2005). Retrieved from: https://www.ncbi.nlm.nih.gov/pubmed/15759309
3.) Sven Bogdan, Klämbt, Epidermal growth factor receptor signaling (2001), pg 3. Retrieved from: https://www.cell.com/current-biology/comments/S0960-9822(01)00167-1
4.) Thomas, Weihua, Rethink of EGFR in Cancer With Its Kinase Independent Function on Board (August, 2019). Retrieved from: https://www.frontiersin.org/articles/10.3389/fonc.2019.00800/full
5.) Tian, Shi, et. al (2018). Different subtypes of EGFR exon19 mutation can affect prognosis of patients with non-small cell lung adenocarcinoma [Pubmed Article]. Retrieved from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6211626/
6.) Tabeta, Hoebe, et al, Velvet, a Dominant Egfr Mutation That Causes Wavy Hair and Defective Eyelid Development in Mice (2003). Retrieved from: https://pdfs.semanticscholar.org/7eee/c89406e69357fa8ccf3874696448de616699.pdf?_ga=2.21516347.1433771477.1588236341-1015871252.1588236341
7.) Ohashi, Rai, et al., Induction of lung adenocarcinoma in transgenic mice expressing activated EGFR driven by the SP‐C promoter (2008). Retrieved from: https://www.ncbi.nlm.nih.gov/pubmed/18564139
8.) Martin, Chemical approaches for profiling dynamic palmitoylation (2013). Retrieved from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3693551/
9.) Cancer Treatment Centers of America, Lung cancer types, pg 2. Retrieved from: https://www.cancercenter.com/cancer-types/lung-cancer/types
10.) Uramoto, Iwata, et. al (2010). Epithelial–Mesenchymal Transition in EGFR-TKI Acquired Resistant Lung Adenocarcinoma [Pubmed Article]. Retrieved from: https://www.ncbi.nlm.nih.gov/pubmed/20682976
11.) Sawabata, Circulating tumor cells in lung cancer: cluster circulating tumor cells as hybrid epithelial-mesenchymal transition/mesenchymal-epithelial transition (2017). Retrieved from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5723813/
12.) Lin, Wang, et al., EGFR-TKI resistance in NSCLC patients: mechanisms and strategies (2014). Retrieved from: https://www.nature.com/articles/onc2012158
13.) Singh, Behrens, et al., Phosphorylation of MUC1 by Met Modulates Interaction with p53 and MMP1 Expression (2008). Retrieved from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4163608/