What is Gene Ontology?
Gene ontology is an aspect of bioinformatics that organizes information in a comprehensive way. For example, genetic information is graphically represented to explicitly show the relationship between each set of data about a specific gene [1]. In this case, EGFR can be studied with the Gene Ontology Database and Uniprot, which are corpuses of information relating gene ontology terms. In the image bellow, the three main topics are outlined from a similar online tool, Quickgo: The biological process, molecular function and cellular components are all important parts of genetic prevalence over diseases.
Can ontological knowledge bases yield information on EGFRs function in LADC? Is EGFR a motivator of LADC phenotypes, such as cell proliferation and invasive phenotypes? Compiling information from these corpuses and investigating the relationship between genes can allow new inferences to be made.
Results
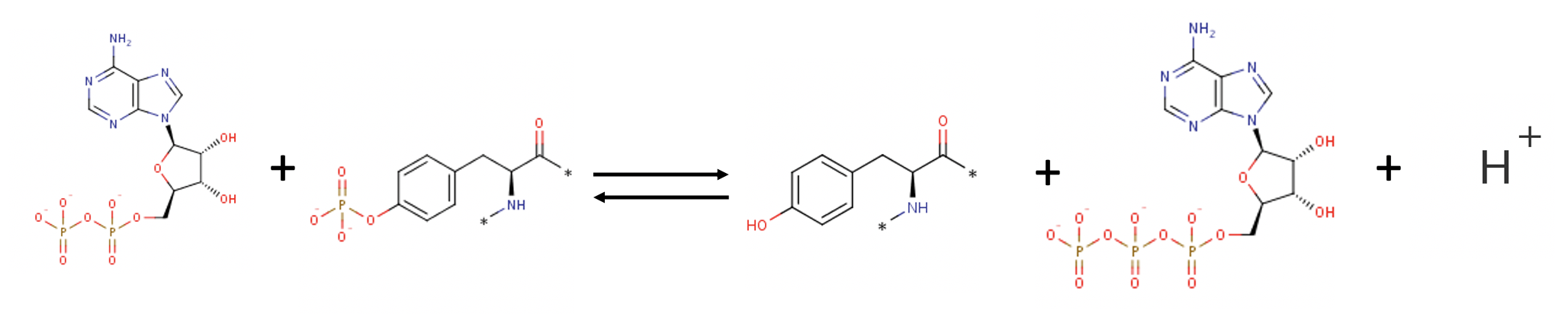
Molecular function- The molecular activity of EGFR is mostly characterized by its Tyrosine Kinase Activity [2]. It also plays a role in intercellular signaling upon ligand binding. Autophosphorylation, a phosphorylation event that occurs after dimerization, results in binding with intracellular molecules. These mechanisms contribute to cellular function and biological roles of EGFR.
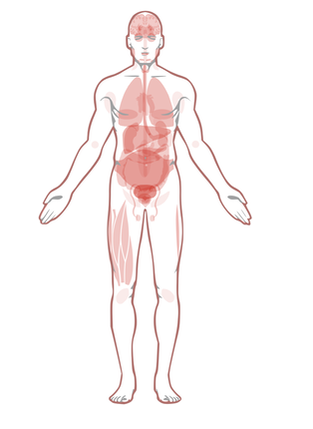
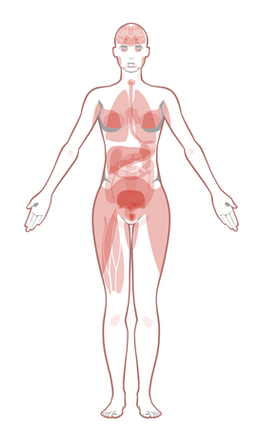
Biological Process- EGFR is expressed in epithelial tissue throughout the body. In addition, EGFR has been implicated in several types of cancers throughout the body including colon, cervical and breast cancer. This is concurrent with the results from Protein Atlas shown bellow.
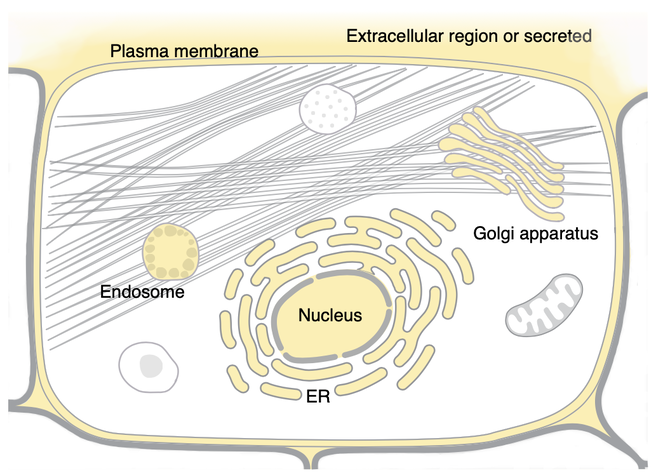
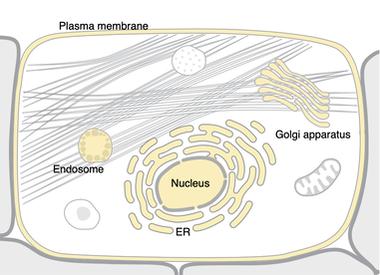
Cellular Component- The cellular components and complexes in which the EGFR gene can carry out its role as a transmembrane receptor is important. In humans, EGFR can be translocated to the Golgi apparatus and the Endoplasmic reticulum, which may be relevant to its molecular function.
Discussion
The EGFR gene has been implicated in the cell cycle, transcription, calcium transport and cell signaling as indicated by Uniprot. In the context of LADC, cell survival, invasion and metastasis are all important. For example, Gene ontology can enrich gene expression data from cancer patients. Visit the transcriptonomics page for an example.
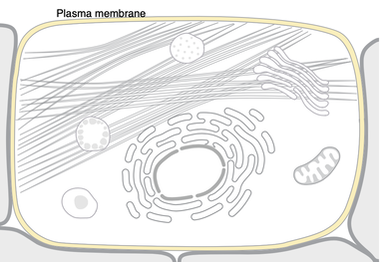
Studying EGFR may be more useful in models that incorporate these cellular, biological and molecular processes. Mice, which share similar cellular presence, may be better to study LADC. As shown in the above images from Uniprot, Zebrafish EGFR seems to only localize to the plasma membrane rather than multiple organelles. In other words, EGFR transport may be important to cancer progression. For example, TKI driven stress on Endoplasmic Reticulum has been studied in intestinal epigeal cells [3].
References
1.) David P Hill, Barry Smith, et al., Gene Ontology annotations: what they mean and where they come from (). Retrieved from:https://bmcbioinformatics.biomedcentral.com/articles/10.1186/1471-2105-9-S5-S2bmcbioinformatics.biomedcentral.com/articles/10.1186/1471-2105-9-S5-S2
2.) Zanetti-Domingues, Korovesis, et al, 2018, The architecture of EGFR’s basal complexes reveals autoinhibition mechanisms in dimers and oligomers. Retrieved From: https://www.nature.com/articles/s41467-018-06632-0
3.) Qiang, Xu, et al., EGFR inhibitor-driven endoplasmic reticulum stress-mediated injury on intestinal epithelial cells Author links open overlay (2014). Retrieved from: https://www.ncbi.nlm.nih.gov/pubmed/25445223
2.) Zanetti-Domingues, Korovesis, et al, 2018, The architecture of EGFR’s basal complexes reveals autoinhibition mechanisms in dimers and oligomers. Retrieved From: https://www.nature.com/articles/s41467-018-06632-0
3.) Qiang, Xu, et al., EGFR inhibitor-driven endoplasmic reticulum stress-mediated injury on intestinal epithelial cells Author links open overlay (2014). Retrieved from: https://www.ncbi.nlm.nih.gov/pubmed/25445223