What are protein post-modifications?
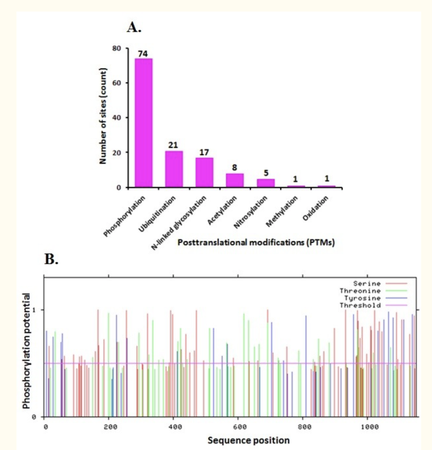
Protein post-translational modifications increase the diversity of the proteome by the covalent addition, proteolytic cleavage of regulatory elements, or degradation [1]. These modifications include phosphorylation, ubiquitination, glycosylation, nitrosylation and methylation amongst many others which influence normal cell biology and pathogenesis. Therefore, identifying and understanding post-translational modifactions is critical in the study of genetics and disease treatment.
|
There are several post-translational modifications that may play a role in LADC. On the left [2], the sequence position is indicated in relation to phosphorylation potential. The C-terminal end of the protein seems to have the highest potential. However, there are also other protein post-modifications that may be important for EGFR's role in cellular processes. Ubiquitation and glycosylation are amongst some important modifications. Ubiquitation, for example can result in protein specific signaling or transport.
|
Using online resources to predict post-modifications can be compared to literature on this gene to deduce important characteristics of EGFR. NetPhos and Nexus were used to accomplish this.
Results
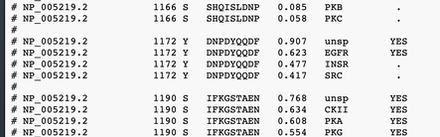
The epidermal growth factor receptor contains amino acids that may be phsop[horylated. NetPhos is a web service that searchers for amino acids that are in the right sequential context for phosphorylation sites.
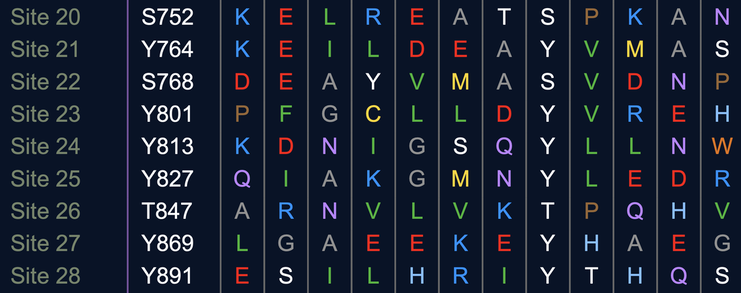
The image bellow shows another prediction of possible phosphorylation sites from OncoNet. Site 26 is proximalto the 858 position, which is implicated in cancer progression. It seems like there may be modifications near this position that may be important.
Discussion
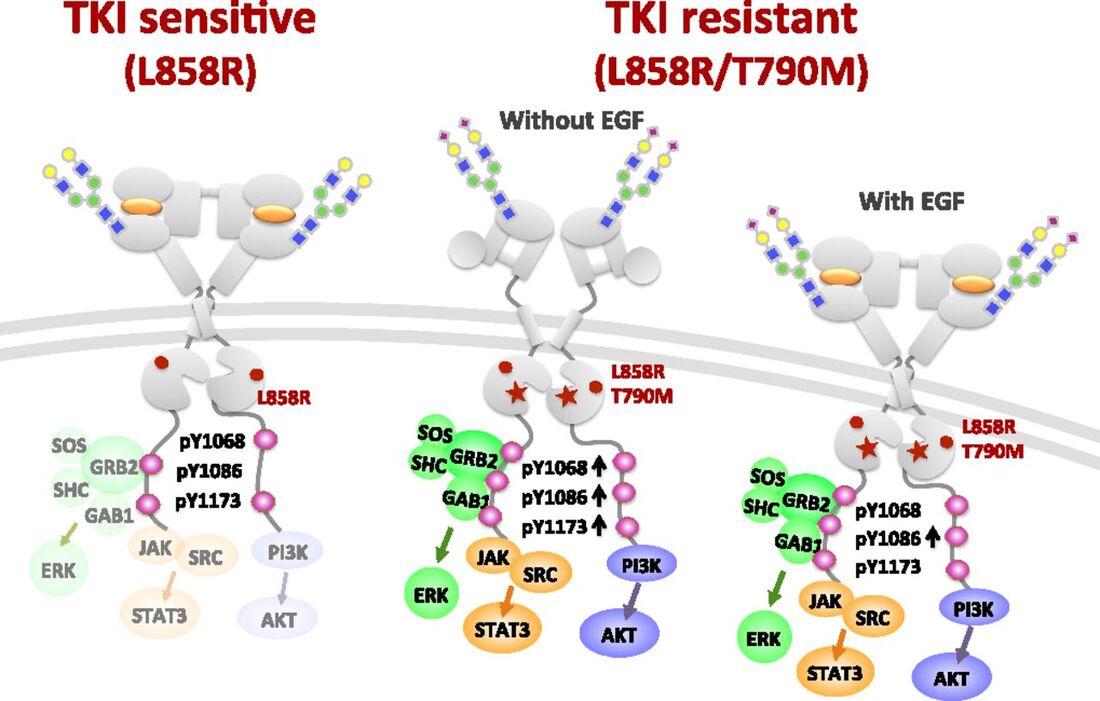
Comparing these results with knowledge of EGFR's catalytic activity as well as specific binders may be relevant to anti-cancer drugs. For example, TKI resistance changes depending on the binding to EGFR post-modifications as shown bellow.
References
1.) Mary Johnson, Protein Modification. Retrieved From: https://www.labome.com/method/Protein-Modification.html
2.) Douglas Arenberg, MD, Radon: A Modifiable Lung Cancer Risk Factor, 2018. Retrieved from:https://www.lungcancernews.org/2018/02/19/radon-a-modifiable-lung-cancer-risk-factor/
3.) Pirker, Hirsch, et al., Consensus for EGFR Mutation Testing in Non-small Cell Lung Cancer: Results from a European Workshop, 2010. Retrieved from: https://www.sciencedirect.com/science/article/pii/S1556086415318220
4.) Purba, Saita, et al., Activation of the EGF Receptor by Ligand Binding and Oncogenic Mutations: The “Rotation Model”, 2017. Retrieved from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5492017/
2.) Douglas Arenberg, MD, Radon: A Modifiable Lung Cancer Risk Factor, 2018. Retrieved from:https://www.lungcancernews.org/2018/02/19/radon-a-modifiable-lung-cancer-risk-factor/
3.) Pirker, Hirsch, et al., Consensus for EGFR Mutation Testing in Non-small Cell Lung Cancer: Results from a European Workshop, 2010. Retrieved from: https://www.sciencedirect.com/science/article/pii/S1556086415318220
4.) Purba, Saita, et al., Activation of the EGF Receptor by Ligand Binding and Oncogenic Mutations: The “Rotation Model”, 2017. Retrieved from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5492017/