What are protein interaction networks?
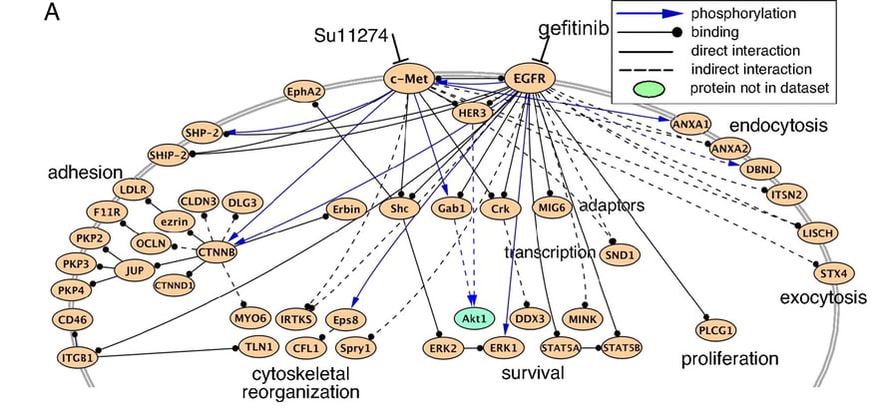
A genetic interaction between genes plays a major role in controlling cellular processes. In the last decade, genome scale studies of quantitative genetic interactions raise questions regarding the functional relationship of genetic and molecular interaction networks and the evolutionary conservation of certain molecular players [1]. As shown in the image bellow, genetic interaction encompasses direct and indirect interaction as well as phosphorylation and binding.
Also illustrated in the image is the role that EGFR interactions have over the cell. TKIs such as gefinib may affect these pathways, which is why it's important to understand the prevalence of interaction networks. Services like the String database can generate interaction diagrams that can colorize dozens of interactions.
Results
Fully active EGFR occurs in cells as a heterodimer. An EGFR monomer comprises of an N-terminal ligand-binding extracellular domain and an intracellular module connected by a single-pass transmembrane helix. The extracellular segment is made up of four domains (I–IV) which connects a small transmembrane segment and a tyrosine kinase domain (TKD). Ligand binding stabilizes the dimer conformation thereby causing an allosteric interaction, allowing Grb2 binding to the intercellular domain (Yamazaki, Zaal, et al., 2002, pg.2).
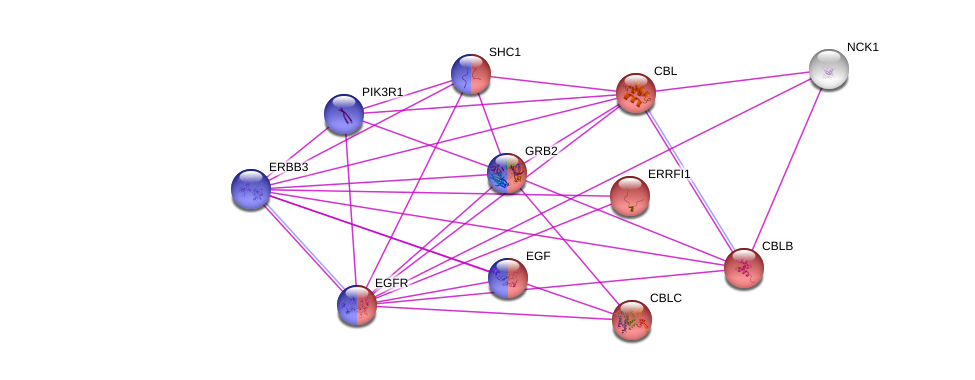
Using the String database, several interaction networks showed that tyrosine kinase function is related to other genes involved in both similar and different function.
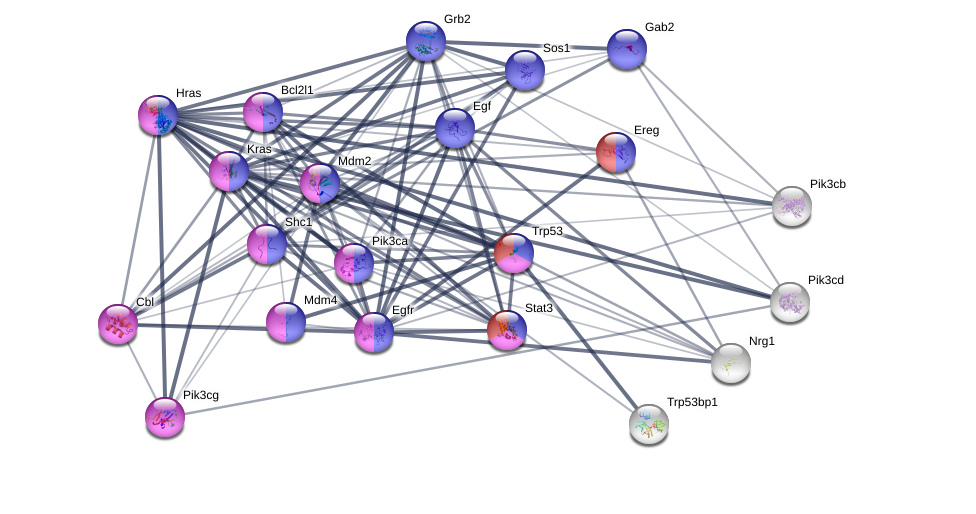
String searches can also be generated based on organism.Bellow is an addition of genes in mice as well as gene ontology parameters, which was transncritiojnn regulation (purple).
Discussion
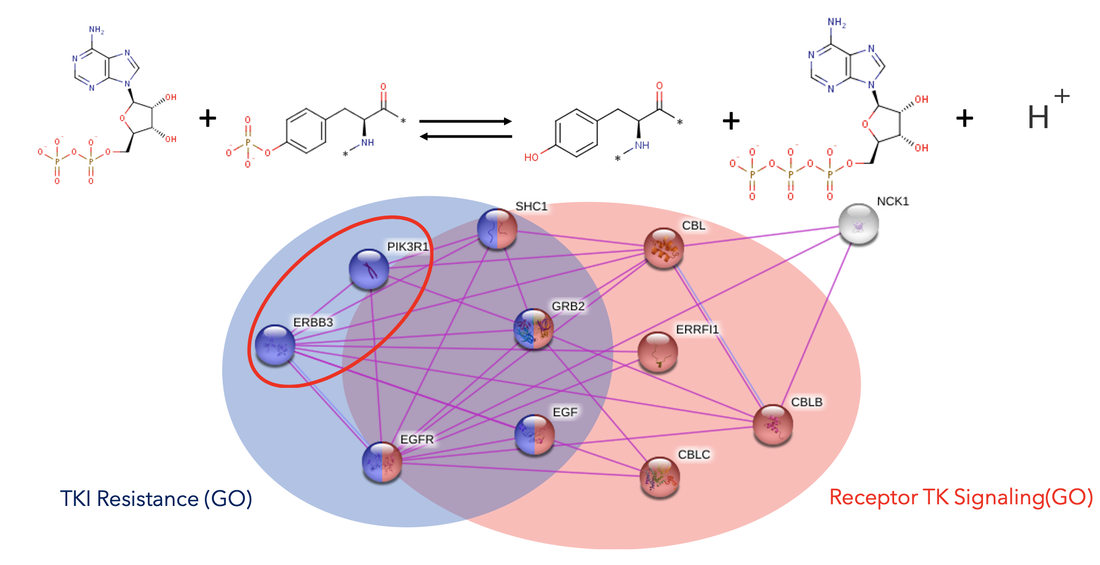
Comparing information about gene ontology terms such as tyrosine kinases can yield important information about receptor function. Although tyrosine kinase function seems to be prevalent over EGFR signaling, there seems to be genes that might function independently of tyrosine kinase activity [2].
References
1.) Janusz Dutkowski, Michael Kramer, A gene ontology inferred from molecular networks (2013). Retrieved From: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3654867/
2.) Thomas, Weihua, Rethink of EGFR in Cancer With Its Kinase Independent Function on Board (August, 2019). Retrieved from: https://www.frontiersin.org/articles/10.3389/fonc.2019.00800/full
3.) Pirker, Hirsch, et al., Consensus for EGFR Mutation Testing in Non-small Cell Lung Cancer: Results from a European Workshop, 2010. Retrieved from: https://www.sciencedirect.com/science/article/pii/S1556086415318220
2.) Thomas, Weihua, Rethink of EGFR in Cancer With Its Kinase Independent Function on Board (August, 2019). Retrieved from: https://www.frontiersin.org/articles/10.3389/fonc.2019.00800/full
3.) Pirker, Hirsch, et al., Consensus for EGFR Mutation Testing in Non-small Cell Lung Cancer: Results from a European Workshop, 2010. Retrieved from: https://www.sciencedirect.com/science/article/pii/S1556086415318220